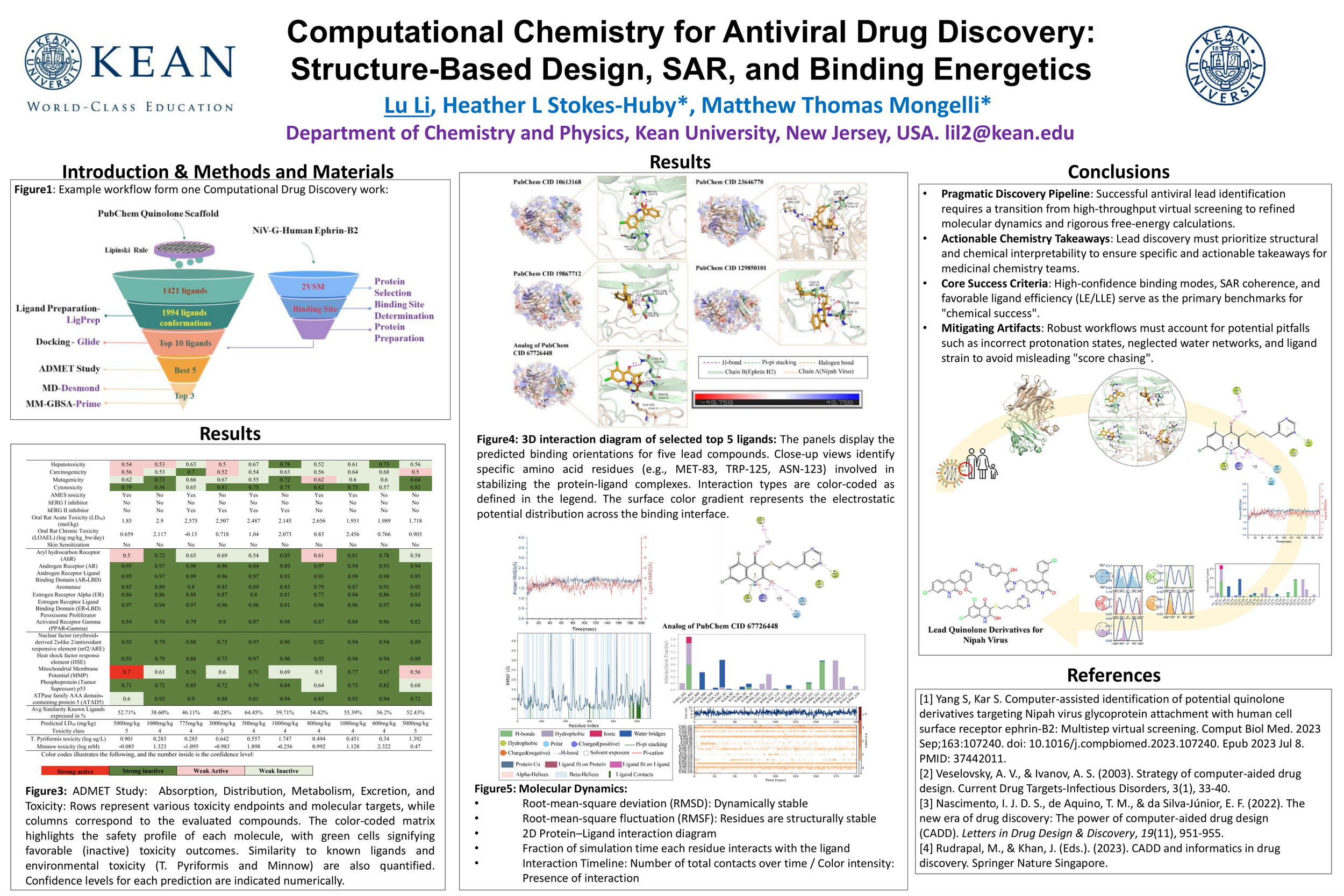
Computational Chemistry for Antiviral Drug Discovery: Structure-Based Design, SAR, and Binding Energetics
Lu Li
Co-Presenters: Individual Presentation
College: Hennings College of Science Mathematics and Technology
Major: BS.BIO/CELL/MOLEC
Faculty Research Mentor: Matthew Mongelli, Heather Stokes-Huby
Abstract:
Zika virus (ZIKV) is a mosquito-borne flavivirus associated with congenital Zikasyndrome and neurological complications, yet no approved antiviral therapies are currentlyavailable. The viral nonstructural protein 5 (NS5) is essential for viral replication and immuneevasion and contains two highly conserved enzymatic domains: a methyltransferase (MTase)required for 5′ RNA capping and an RNA-dependent RNA polymerase (RdRp) responsible forviral genome replication. Simultaneous inhibition of both domains represents a promisingantiviral strategy with the potential to enhance efficacy and reduce resistance.In this study, a structure-based virtual screening workflow was applied to identify dual-target NS5 inhibitors. Crystal structures of ZIKV MTase (PDB 5WXB) and RdRp (PDB 5U04)were obtained from the Protein Data Bank and prepared using Schrödinger. Binding sites wereidentified using SiteMap, and docking grids were generated accordingly. A ChemDiv compoundlibrary consisting of 53,121 small molecules, supplemented with 10 literature-reported controlcompounds, was processed with LigPrep to generate protonation states and stereoisomers atphysiological pH.Virtual screening was conducted using Glide in a hierarchical manner, progressing fromhigh-throughput virtual screening (HTVS) to extra-precision (XP) docking. Compounds weredefined as dual-target candidates if they achieved docking scores below −6.0 kcal/mol againstboth MTase and RdRp. Of the initial library, 283 compounds advanced to XP docking, and 12met the dual-target selection criteria. These candidates were further evaluated using ProTox-IIIto predict toxicity endpoints, including hepatotoxicity, mutagenicity, and carcinogenicity.Based on combined docking performance and predicted safety profiles, five compounds(CE03-6244, CE04-4915, CE04-6933, CE05-6995, and CE05-9535) were prioritized for 500 nsmolecular dynamics simulations. Protein–ligand interaction analyses revealed recurrenthydrogen bonding and electrostatic interactions with key active-site residues in both MTase andRdRp, supporting stable dual-domain binding. Overall, this computational study identifiespromising dual-target NS5 inhibitor candidates and provides a foundation for further dynamicsimulations and experimental validation.